- Record: found
- Abstract: found
- Article: found
Divide and Conquer: High Resolution Structural Information on TRP Channel Fragments
research-article
Read this article at
There is no author summary for this article yet. Authors can add summaries to their articles on ScienceOpen to make them more accessible to a non-specialist audience.
Abstract
Understanding how proteins facilitate signaling and substrate transport across biological
membranes is an important frontier of structural biology. Membrane proteins are the
doors and windows of cells: many membrane proteins are gates of entry into or exit
from cells or cellular compartments, and others allow cells to sense their environment.
One important multifunctional family of membrane proteins is the transient receptor
potential (TRP) family of ion channels. TRP channels have recently been the subject
of multiple structural analyses, both low resolution electron microscopy studies (reviewed
by Moiseenkova-Bell and Wensel in this issue [p. 239]) and the divide and conquer
approach of determining high resolution crystal structures of channel fragments, reviewed
here.
Introduction
TRP channels form a large family of cation channels that can be activated by diverse
signals, including chemical ligands and/or temperature or mechanical stimuli (Ramsey
et al., 2006; Venkatachalam and Montell, 2007). Most TRP channels are also modulated
by various intracellular signals, including calcium, phosphoinositides, and other
lipid metabolites. TRP channels are mostly found in the animal kingdom (organisms
with a nervous system), consistent with their prominent role in sensory perception.
They are distributed into seven subfamilies according to sequence and function (Montell,
2005): TRPA (ankyrin), TRPC (canonical), TRPM (melastatin), TRPML (mucolipin), TRPN
(NOMPC), TRPP (polycystin), and TRPV (vanilloid). Of note, TRPN channels are found
in most animal genomes but excluded from mammalian ones.
The structural biology of ion channels is an important and expanding research endeavor.
Mechanistic understanding of ion channel function is central to our understanding
of neurobiology and many other physiological processes. Furthermore, ion channels
are important targets for drug development. With the rapidly increasing number of
structures of ion channels and their fragments (Minor, 2007), including structural
studies of TRP channels (Gaudet, 2008b), there is an opportunity to leverage this
structural information in studies of TRP channel function and physiology. TRP channel
biologists and physiologists may want to brush up on structural biology approaches,
and an excellent starting point is a recent primer on structural biology for neurobiologists
(Minor, 2007). Conversely, structural biologists benefit from integrating knowledge
on TRP channels and general channel physiology in planning their experiments. TRP
channels are challenging structural biology targets, and the more that is known about
their molecular properties, the more likely we will be to succeed in obtaining valuable
three-dimensional structures.
In structural biology, the aim is to understand proteins at several levels: What are
their structural and functional modules? How is the modular architecture integrated
to drive their molecular mechanisms? How are these proteins incorporated into larger
assemblies? How do these assemblies regulate protein function in a cellular context?
Furthermore, the integration of structural and physiological approaches enables us
to advance from static three-dimensional structures to the description of dynamic
processes like conformational changes and ligand interactions.
Determining the high resolution structure of complete TRP channels remains a major
challenge. One alternative and complementary strategy is to divide and conquer: determine
crystal structures of isolated domains of TRP channels. The resulting information
can then be pieced together and integrated with biochemical and physiological data
to advance our understanding of TRP channel function. Below, I describe how the divide
and conquer approach can be implemented, illustrate some recent results obtained with
fragments of TRPV and TRPM channels, and pinpoint some challenges that lie ahead in
moving from this piecemeal approach to the ultimate goal of obtaining a full molecular-level
description of TRP channel function.
TRP Channels as Modular Proteins
TRP channel subunits are rather large, ranging from ∼700 to more than 2,000 amino
acid residues, and have six membrane–spanning segments with an extended pore loop
between the fifth and sixth segment. This transmembrane domain arrangement is homologous
to that of other ion channels in a large superfamily that includes voltage-gated calcium
channels and Shaker potassium channels (Venkatachalam and Montell, 2007). All members
of this superfamily are believed to assemble as tetramers of the six-segment transmembrane
domain, with a central ion permeation path. Most if not all TRP proteins can homotetramerize
to form functional channels, and several also have the ability to heterotetramerize,
thus increasing the permutations of possible functional units (for a recent review
see Schaefer, 2005).
The transmembrane domain of TRP proteins spans ∼300 residues and is connected at the
N and C termini to large intracellular regions containing protein-interaction and
regulatory motifs with distinctive features for each TRP subfamily (for a recent review
see Gaudet, 2006). For instance, ankyrin repeats are ubiquitous ligand-interaction
motifs that are found in the N-terminal cytosolic region of TRPC, TRPV, TRPA, and
TRPN channels. As an example, Fig. 1 illustrates the relationship between the ankyrin
repeats and other regions of TRPV channels. As a contrasting example, the overall
domain structure of a TRPM channel is depicted in Fig. 2. TRPM channels do not have
ankyrin repeats but instead have a large, ∼700-residue N-terminal intracellular region
homologous only to other TRPM channels. C-terminal to the transmembrane domain, TRPM
channels have a coiled-coil domain. In some TRPM channels, an enzymatic domain then
follows the coiled-coil domain: TRPM6 and TRPM7 have an α-kinase domain (Nadler et
al., 2001; Runnels et al., 2001), and TRPM2 has a NUDIX domain that interacts with
ADP-ribose nucleotides (Perraud et al., 2001).
Figure 1.
The ankyrin repeats of TRPV channels. Diagram shows the topology of TRPV channels
with the relative position of the ankyrin repeats illustrated with the structure of
the TRPV1 ARD. Only two of the four subunits are shown for clarity in yellow and green,
respectively; the subunits in front and in back of the plane of the page are omitted.
The transmembrane domains are illustrated using the homologous structure of the Shaker
potassium channel (Long et al., 2005). The N- and C-terminal segments of unknown structure
are depicted with shapes that approximate their relative size. ATP and ATP-interacting
side chains are shown as sticks and colored according to atom type, and a transparent
surface representation highlights the surface complementarity of the ATP and its binding
site. The approximate size of those protein segments in numbers of amino acid (aa)
residues is indicated for the green subunit. The transmembrane and ARDs of TRPV channels
are each ∼250 amino acid residues. TRPV subunits typically are ∼800-residues long.
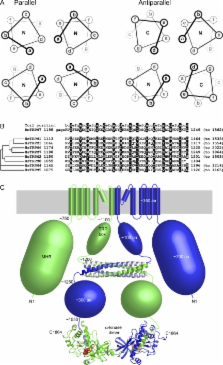
Figure 2.
The coiled-coils of TRPM channels. (A) Helical wheel representation of parallel (left)
and antiparallel (right) tetrameric coiled-coils. The “N” or “C” in each wheel indicates
whether the N or C terminus, respectively, of the α-helix points toward the viewer.
Darker lines are in front and lighter ones are in the back. In both coiled-coils,
the a and d residues of the heptad repeats form the core, whereas the e and g residues
form peripheral interactions. However, the details of the interactions are different.
In a parallel coiled-coil, each layer of hydrophobic interactions consists of either
four a or four d residues. In contrast, each antiparallel layer consists of two d
and two a residues. (B) Sequence alignment of the predicted coiled-coil sequences
of human TRPM channels. The sequence of the rat TRPM7 coiled-coil, for which the structure
is available, is also included at the top. a and d position residues are shaded. The
dendrogram was generated by ClustalW using an alignment of whole TRPM sequences. (C)
Diagram of TRPM7 displaying the available structural information. Similarly to Fig.
1, only two of the four channel subunits are illustrated for clarity (green and blue,
respectively), except for the coiled-coil structure where all four strands are shown.
Note that the transmembrane domain would have fourfold rotational symmetry perpendicular
to the membrane (gray shading), whereas the coiled-coil and α-kinase domains only
have twofold symmetry. Shape sizes approximate the number of residues in each region,
and the size (in number of amino acid residues [aa]) is indicated for the blue subunit.
The approximate boundaries, in residue numbers, of different domains are also indicated.
How can this modular domain structure of TRP channels be leveraged in structural biology?
A fundamental element of a successful divide and conquer approach to protein structure
determination is to properly identify the boundaries of TRP channel domains to enable
the expression and purification of these domains in isolation. The word “domain” is
often used rather loosely by nonstructural biologists (and sometimes even structural
biologists) to describe any fragment a protein—often, the terms “segment,” “region,”
or “motif” would be more appropriate. The structural biology definition of a domain
is a compact globular structure that can fold autonomously and originates from early
structural studies of immunoglobulins (Wetlaufer, 1973). This definition implies that
(1) a domain is large enough to have a unique three-dimensional fold, and (2) a domain
can often fold on its own in isolation from the rest of the protein. This second point
is the key to a divide and conquer approach because by identifying the proper domain
boundaries of a region of interest, it can then be isolated for structural studies.
Furthermore, a protein domain is a self-contained unit that can interact with other
molecules or other parts of the protein. In a divide and conquer approach, one can
therefore still obtain information about relevant regulatory interactions by determining
structures of domains with their respective ligands. From a genomics perspective,
a domain can also evolve new functionalities and be swapped in and out of genes during
evolution by duplication or deletions (Moore et al., 2008).
The divide and conquer approach to TRP channel structural biology has thus far yielded
structures of two different types of domains, ankyrin repeats from TRPV channels and
a coiled-coil from a TRPM channel. Both types of domains are found not just in TRP
channels, but also in many other protein families. At first glance, that might lead
one to think that the resulting structures are old news; after all, there are many
published structures of ankyrin repeats and coiled-coils (for recent reviews see Gaudet,
2008a and Grigoryan and Keating, 2008). But in both cases, structures of some of their
representatives in TRP channels have yielded surprises. The next two sections describe
the lessons we have thus far learned from structures of TRP channel ankyrin repeats
and coiled-coils.
Lessons from Ankyrin Repeats
Mammalian TRPV channels are divided into two subgroups: TRPV1 through TRPV4 mediate
responses to many sensory stimuli, including heat, low pH, neuropeptides and chemical
ligands, whereas TRPV5 and TRPV6 are expressed in the kidney and gut, respectively,
and are involved in calcium homeostasis (Venkatachalam and Montell, 2007). Several
TRPV channels are polymodal detectors. For example, TRPV1 is activated not only by
noxious heat, but also by capsaicin and low extracellular pH. The intracellular N-terminal
region of TRPV proteins contains six ankyrin repeats, short sequence motifs often
involved in protein–protein interactions (Gaudet, 2008a). The isolated TRPV ankyrin
repeat domains (ARDs) do not oligomerize, suggesting that the ARDs interact with regulatory
factors instead (Phelps et al., 2008).
Ankyrin repeat sequences span ∼33 residues and fold into a structural motif consisting
of two α-helices folding back onto each other to form a helical hairpin, followed
by a long hairpin loop that extends roughly perpendicular to the helical axes. Multiple
such structural motifs are stacked side by side with their helices nearly parallel
to each other to form an ARD, with the number of repeats ranging from 3 to >30 (Gaudet,
2008a). The structures of several TRPV ARDs have now been published: rat TRPV1 (Lishko
et al., 2007), both rat (Jin et al., 2006) and human TRPV2 (McCleverty et al., 2006),
and mouse TRPV6 (Phelps et al., 2008), and their folds are very similar to each other,
consistent with their sequence homology. The TRPV ARDs have six ankyrin repeat motifs,
with atypical long finger loops and a pronounced twist between the fourth and fifth
repeat, such that the helices of repeats 1–4 and 5–6 are no longer nearly parallel
to each other (Fig. 1). Both the long loops and the unusual twist break the regularity
of the repeats, giving the TRPV ARDs a unique shape. Because both the long loops and
the inter-repeat twist are caused by residues that diverge from the ankyrin repeat
sequence consensus but are conserved in TRPV proteins, it is expected that this unique
shape will be observed in all TRPVs (Phelps et al., 2008).
The unique shape of the TRPV ARDs, while of interest to structural biologists investigating
repeat proteins and protein folding and design, is not particularly informative about
the role of the ARD in TRPV channel function. However, when hundreds of chemicals
were screened to optimize the TRPV1-ARD crystallization conditions, it was observed
that the presence of ATP altered the crystal shape, likely by changing the packing
interactions between protein molecules. This new crystal form diffracted to higher
resolution, allowing structure determination and refinement. The resulting electron
density map indicated that an ATP molecule was indeed bound to the TRPV1-ARD (Fig.
1) (Lishko et al., 2007) on the concave surface that is typically occupied by ligand
in ARD–ligand complexes (Gaudet, 2008a). Biochemical assays demonstrated that both
ATP and calcium calmodulin bind to this same binding surface in a competitive manner—the
binding of one excludes the binding of the other. Another clue that the TRPV1-ARD
interaction with ATP is physiologically relevant is that it is conserved in the chicken
homologue (Phelps et al., 2007), indicating that it is better conserved than capsaicin
sensitivity because chicken TRPV1 is insensitive to capsaicin (Jordt and Julius, 2002).
In electrophysiology experiments, intracellular ATP prevented desensitization to repeated
applications of capsaicin, whereas calcium calmodulin plays an opposing role and was
required for desensitization (Lishko et al., 2007). The accumulated data lead to a
model for the calcium-dependent regulation of TRPV1 via the competitive interactions
of ATP and calmodulin at the N-terminal binding site. In summary, the crystallographic
determination of the TRPV1-ARD structure has lead to the fortuitous discovery of a
regulation mechanism for TRPV1. It will be interesting to see whether this mechanism
is conserved in other TRPV ion channels.
Ankyrin repeats are also found in other TRP channels, including TRPA, TRPN, and TRPC
channels. The TRPC and TRPV channels have few repeats and irregular sequences (Phelps
et al., 2007, 2008), whereas TRPA and TRPN channels have many regular repeats (for
a recent review see Gaudet, 2008a). As was done in the case of TRPV channels, the
abundant information on ankyrin repeats from both natural and designed proteins can
be leveraged to study the role of ankyrin repeats in other TRP channels. Of particular
interest is TRPA1, which transduces pain signals in response to irritants like mustard
oil (Bandell et al., 2004). Irritants covalently attach to the thiol group of several
cysteines in TRPA1’s 17 ankyrin repeats to activate the channel (Hinman et al., 2006;
Macpherson et al., 2007), and structures of the ankyrin repeats will be useful to
decipher how this chemical modification can lead to channel opening. Allicin, a compound
found in garlic, activates TRPV1 through the chemical modification of a single cysteine,
C157, in the ARD of TRPV1 (Salazar et al., 2008). Cysteine 157 is buried in the protein
core between repeats 1 and 2, implying that its chemical modification requires a fairly
large conformational change (Gaudet, 2008b; Salazar et al., 2008). Similarly to the
TRPA channels, TRPN channels have large numbers of ankyrin repeats with sequences
very close to ankyrin repeat motif consensus, although little is known about the biological
roles of these repeats. TRPC channels have few repeats (likely four or five), which
have weak similarity to ankyrin repeat consensus. The structure of TRPC channel ankyrin
repeats is therefore likely to have some unusual kinks and loops, as was observed
in TRPV channels.
Lessons from Coiled-Coils
The TRPM channels have coiled-coil domains in their C-terminal cytosolic region. Biophysical
studies (Tsuruda et al., 2006) have validated the existence of these coiled-coils
in all but one of the eight mammalian TRPM channels (TRPM1 was not validated in this
study) and demonstrated that these coiled-coils can form homotetrameric assemblies,
which is consistent with the expected tetrameric state of functional TRPM channels.
Coiled-coils are protein interaction and assembly motifs forming α-helices that zip
up together in a helical coil conformation (for a recent review see Grigoryan and
Keating, 2008). Coiled-coils are found in many protein families, including transcription
factors, cellular and viral membrane fusion proteins, and ion channels. Coiled-coil
motifs are identified in protein sequences by their characteristic recurring pattern
of aliphatic residues alternating every third then fourth residue to form seven-residue
repeats. The sequence patterns are a reflection of the regularity of three-dimensional
coiled-coil structures (Fig. 2 A). Within each repeat of seven amino acids, routinely
labeled a through g, residues a and d are usually aliphatic and form the internal
core of the coiled-coil. Residues e and g are generally polar or charged and interact
with each other across strands, often dictating the specificity of assembly through
electrostatic interactions. Residues b, c, and f tend to lie on the outside surface
and have less influence on coiled-coil interactions.
Although the above description may suggest that coiled-coil structures are predictable,
this is currently not the case (Grigoryan and Keating, 2008). Coiled-coil structures
have been observed that contain anywhere between two and seven helical strands, and
strands can associate in either parallel or antiparallel orientations. The number
and orientation of the strands in a coiled-coil assembly cannot be predicted. Small
variations in coiled-coil sequences, as little as one residue, can change the observed
assembly mode (Grigoryan and Keating, 2008). Furthermore, some strands preferentially
form homo-oligomers, whereas others form specific hetero-oligomers. Repeat residues
e and g play important roles in dictating specificity, but in ways that cannot yet
be predicted easily. In summary, the identification of coiled-coil repeats can enable
the prediction of which residues are most likely to mediate affinity (a and d) and
specificity (e and g), and which residues may have little influence on assembly (b,
c, and f). But the nature of the resulting assembly cannot be predicted with certainty.
TRPM6 and TRPM7 are two closely related TRPM family members that are important in
magnesium uptake and homeostasis (Schlingmann et al., 2007). The crystal structure
of the TRPM7 coiled-coil was recently determined to high resolution (Fujiwara and
Minor, 2008). It forms a homotetrameric coiled-coil, consistent with the predicted
tetrameric functional channel. But surprisingly, it is an antiparallel tetrameric
coiled-coil, with two strands going in one direction and two strands going in the
opposite direction (Fig. 2 C). This is striking because it breaks the fourfold rotational
symmetry that is expected for the transmembrane domain of the TRPM7 channel, with
the four subunits related by 90° rotations around an axis perpendicular to the plane
of the membrane. The antiparallel topology was confirmed in solution using cross-linking
experiments (Fujiwara and Minor, 2008). It will be important to further confirm that
the antiparallel topology observed for the isolated coiled-coil domain is also present
in intact TRPM7 channels, although sequence analyses do strongly support an antiparallel
topology for the TRPM7 coiled-coil and closely related TRPM channels (Fujiwara and
Minor, 2008).
The TRPM7 coiled-coil is followed by an atypical α-kinase domain. The structure of
that kinase domain, determined in 2001 (Yamaguchi et al., 2001), showed a domain-swapped
dimer where the two subunits are held together by an exchange of their N-terminal
helices (Fig. 2 C). The ∼80-Å distance between the C termini of two antiparallel strands
of the TRPM7 coiled-coil structure matches well the ∼90-Å distance between the N termini
of one kinase dimer. Therefore, by breaking the fourfold symmetry of the transmembrane
domain, the antiparallel coiled-coil may allow two kinase dimers to exist side by
side. This observation lends further support to the physiological relevance of the
unexpected oligomer symmetry observed in both structures. It also prompts a word of
caution regarding electron microscopy analyses of TRP channel structures. Symmetry
averaging is routinely used to improve the signal to noise when analyzing electron
microscopy images. However, one can no longer assume that a TRP channel obeys fourfold
rotation symmetry throughout the tetrameric assembly.
TRPM6 is the closest TRPM7 homologue that also has a C-terminal α-kinase domain, with
a sequence identity of 53% at the protein level overall and 69% in the coiled-coil
domain. TRPM6 and TRPM7 have been reported to form heterotetramers when coexpressed
in the same cells (Chubanov et al., 2004). It will be interesting to follow up on
the structural studies of the two TRPM7 C-terminal domains with structural and/or
biochemical studies to investigate whether the TRPM6 and TRPM7 coiled-coils can form
heterotetramers, and whether the TRPM6 and TRPM7 kinase domains can form heterodimers.
The results could suggest possible stoichiometries and topologies available to TRPM6/TRPM7
heterotetramers.
The observation that the TRPM7 coiled-coil is antiparallel also brings up the interesting
question of whether all TRPMs will have antiparallel coiled-coils or whether the observed
sequence divergence also encodes structural divergence. The signature pattern of a
coiled-coil is identifiable in all mammalian TRPM proteins (Fig. 2 B) (Fujiwara and
Minor, 2008), but as described above, the topology of a coiled-coil is not readily
predicted and would be worth testing experimentally for each TRPM family member. Because
small changes in a coiled-coil sequence can tilt the balance to favor parallel versus
antiparallel assembly or alter partnering specificity (Grigoryan and Keating, 2008),
the presence of coiled-coil assembly domains might promote rapid evolution and divergence
of subunit assembly and topology in protein families like the TRPM channels.
Inhibitors of coiled-coil assembly have been selected or designed by optimizing affinity
and specificity to compete effectively against the native interactions (Grigoryan
and Keating, 2008). One example is an HIV inhibitor that prevents fusion of the virus
with the cell membrane (Frey et al., 2006). Therefore, the structure of the TRPM7
coiled-coil, and any future TRPM coiled-coil structure, could be used to design inhibitors
of channel assembly and function. Although the isolated TRPM8 coiled-coil had no effect
on TRPM8 function, when it was attached to an accessory TM helix to pre-localize it
to the plasma membrane, it inhibited channel assembly and function (Tsuruda et al.,
2006). This suggests that a molecule that interacts strongly enough to overcome the
high local concentration of the native coiled-coil to disrupt its assembly could be
an effective inhibitor of TRPM8.
Coiled-coils are also predicted in TRPC channels at either or both the N-terminal
intracellular linker between the ankyrin repeats and the transmembrane channel domain
and the C-terminal domain (Lepage and Boulay, 2007; Schindl and Romanin, 2007). These
TRPC coiled-coil regions have yet to be confirmed through biochemical and/or structural
experiments. It will be interesting to see whether future studies of TRPC coiled-coils
will also yield new surprises, considering the interesting recent developments in
the TRPM coiled-coil structural studies.
Future Outlook
To fully understand the molecular basis of TRP channel gating and regulation, high
resolution structures of whole TRP channels will be great assets, whether by electron
microscopy techniques or x-ray crystallography. TRP channel structures will generate
a framework for interpreting biochemical and electrophysiological information accumulated
on these channels. X-ray crystallography of membrane proteins like TRP channels poses
several technical challenges. One technical challenge is to produce large amounts
of detergent-solubilized, biochemically pure TRP channel tetramers. Recent progress
in membrane protein crystallography is encouraging, including structures of vertebrate
ion channels including Shaker channels produced in the yeast Pichia pastoris (Long
et al., 2005) and an ASIC channel produced in baculovirus-infected insect cells (Jasti
et al., 2007). A second technical challenge is obtaining crystals of TRP channels
suitable for structure determination by x-ray crystallography. Crystallization is
still based on trial-and-error methods screening thousands of conditions. Aside from
the typical crystallization solution components (buffering and precipitating agents,
salts, and other chemical additives), membrane proteins require additional screening
with different detergents and/or lipids. Further variables that may prove useful for
TRP channels are the addition of chemical and protein ligands, including agonists,
antagonists, blockers, and other modulators. These ligands have functional effects
on the proteins by changing their conformation, which can in turn influence their
crystal-packing interactions to improve crystal growth.
Detailed mechanistic understanding of TRP channel function will be attained by iterating
structures and functional experiments using physiological assays. Significant advances
have been achieved through studies of channel fragments. But this divide and conquer
approach is ultimately conservative in nature: the long-term goal is to view the channel
as a whole, and although it is perhaps a more risky approach, tackling the structure
of assembled TRP channels will yield information not attainable from the accumulation
of fragmented structures. That is because the divide and conquer approach does not
directly answer the question of how the local information—the conformational state
of the particular fragment under study—is integrated in the context of the whole tetrameric
channel to effect changes in TRP channel function. We can expect that structural information
on TRP channels will continue to emerge, both in fragments and, hopefully, whole channel
structures in the near future. Continued collaboration between physiology and structural
biology will be needed to fully appreciate how complex and elegant TRP channels truly
are.
Related collections
Most cited references27
- Record: found
- Abstract: found
- Article: not found
TRP channels.
Kartik Venkatachalam, Craig Montell (2007)
- Record: found
- Abstract: found
- Article: not found
An introduction to TRP channels.
I. Ramsey, Markus Delling, David Clapham (2006)
- Record: found
- Abstract: found
- Article: not found
Noxious cold ion channel TRPA1 is activated by pungent compounds and bradykinin.
Michael Bandell, Gina M Story, Sun Wook Hwang … (2004)