- Record: found
- Abstract: found
- Article: found
Simultaneous sequencing of genetic and epigenetic bases in DNA

Read this article at
Abstract
DNA comprises molecular information stored in genetic and epigenetic bases, both of which are vital to our understanding of biology. Most DNA sequencing approaches address either genetics or epigenetics and thus capture incomplete information. Methods widely used to detect epigenetic DNA bases fail to capture common C-to-T mutations or distinguish 5-methylcytosine from 5-hydroxymethylcytosine. We present a single base-resolution sequencing methodology that sequences complete genetics and the two most common cytosine modifications in a single workflow. DNA is copied and bases are enzymatically converted. Coupled decoding of bases across the original and copy strand provides a phased digital readout. Methods are demonstrated on human genomic DNA and cell-free DNA from a blood sample of a patient with cancer. The approach is accurate, requires low DNA input and has a simple workflow and analysis pipeline. Simultaneous, phased reading of genetic and epigenetic bases provides a more complete picture of the information stored in genomes and has applications throughout biomedicine.
Abstract
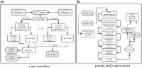
A six-letter sequencing workflow can simultaneously detect genetic and epigenetic bases.
Related collections
Most cited references56
- Record: found
- Abstract: found
- Article: found
fastp: an ultra-fast all-in-one FASTQ preprocessor
- Record: found
- Abstract: found
- Article: found
Twelve years of SAMtools and BCFtools

- Record: found
- Abstract: found
- Article: found