- Record: found
- Abstract: found
- Article: found
Sphenodontian phylogeny and the impact of model choice in Bayesian morphological clock estimates of divergence times and evolutionary rates
Read this article at
Abstract
Background
The vast majority of all life that ever existed on earth is now extinct and several aspects of their evolutionary history can only be assessed by using morphological data from the fossil record. Sphenodontian reptiles are a classic example, having an evolutionary history of at least 230 million years, but currently represented by a single living species ( Sphenodon punctatus). Hence, it is imperative to improve the development and implementation of probabilistic models to estimate evolutionary trees from morphological data (e.g., morphological clocks), which has direct benefits to understanding relationships and evolutionary patterns for both fossil and living species. However, the impact of model choice on morphology-only datasets has been poorly explored.
Results
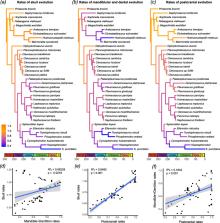
Here, we investigate the impact of a wide array of model choices on the inference of evolutionary trees and macroevolutionary parameters (divergence times and evolutionary rates) using a new data matrix on sphenodontian reptiles. Specifically, we tested different clock models, clock partitioning, taxon sampling strategies, sampling for ancestors, and variations on the fossilized birth-death (FBD) tree model parameters through time. We find a strong impact on divergence times and background evolutionary rates when applying widely utilized approaches, such as allowing for ancestors in the tree and the inappropriate assumption of diversification parameters being constant through time. We compare those results with previous studies on the impact of model choice to molecular data analysis and provide suggestions for improving the implementation of morphological clocks. Optimal model combinations find the radiation of most major lineages of sphenodontians to be in the Triassic and a gradual but continuous drop in morphological rates of evolution across distinct regions of the phenotype throughout the history of the group.
Conclusions
We provide a new hypothesis of sphenodontian classification, along with detailed macroevolutionary patterns in the evolutionary history of the group. Importantly, we provide suggestions to avoid overestimated divergence times and biased parameter estimates using morphological clocks. Partitioning relaxed clocks offers methodological limitations, but those can be at least partially circumvented to reveal a detailed assessment of rates of evolution across the phenotype and tests of evolutionary mosaicism.
Related collections
Most cited references139
- Record: found
- Abstract: found
- Article: found
MrBayes 3.2: Efficient Bayesian Phylogenetic Inference and Model Choice Across a Large Model Space
- Record: found
- Abstract: not found
- Article: not found
Bayes Factors
- Record: found
- Abstract: found
- Article: found