- Record: found
- Abstract: found
- Article: not found
Potent inhibition of diverse Omicron sublineages by SARS-CoV-2 fusion-inhibitory lipopeptides

Read this article at
Abstract
The emergence and rapid spreading of SARS-CoV-2 variants of concern (VOCs) have posed a great challenge to the efficacy of vaccines and therapeutic antibodies, calling for antivirals that can overcome viral evasion. We recently reported that SARS-CoV-2 fusion-inhibitory lipopeptides, IPB02V3 and IPB24, possessed the potent activities against divergent VOCs, including Alpha, Beta, Gamma, Delta, and the initial Omicron strain (B.1.1.529); however, multiple Omicron sublineages have emerged and BA.4/5 is now becoming predominant globally. In this study, we focused on characterizing the functionality of the spike (S) proteins derived from Omicron sublineages and their susceptibility to the inhibition of IPB02V3 and IPB24. We first found that the S proteins of BA.2, BA.2.12.1, BA.3, and BA.4/5 exhibited significantly increased cell fusion capacities compared to BA.1, whereas the pseudoviruses of BA.2.12.1, BA.3, and BA.4/5 had significantly increased infectivity relative to BA.1 or BA.2. Next, we verified that IPB02V3 and IPB24 also maintained their very high potent activities in inhibiting diverse Omicron sublineages, even with enhanced potencies relative to the inhibition on ancestral virus. Moreover, we demonstrated that evolved Omicron mutations in the inhibitor-binding heptad repeat 1 (HR1) site could impair the S protein-driven cell fusogenicity and infectivity, but none of single or combined mutations affected the antiviral activity of IPB02V3 and IPB24. Therefore, we believe that viral fusion inhibitors possess high potential to be developed as effective drugs for fighting SARS-CoV-2 variants including diverse Omicron sublineages.
Related collections
Most cited references44
- Record: found
- Abstract: found
- Article: found
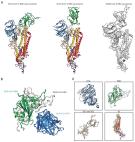
Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation
- Record: found
- Abstract: found
- Article: not found
Structure, Function, and Antigenicity of the SARS-CoV-2 Spike Glycoprotein
- Record: found
- Abstract: found
- Article: not found
