- Record: found
- Abstract: found
- Article: found
The lack of association between ubiquinol‐cytochrome c reductase core protein I ( UQCRC1) variants and Parkinson's disease in an eastern Chinese population
letter
Zhi‐Hao Lin
1 ,
Ran Zheng
1 ,
Yang Ruan
1 ,
Ting Gao
1 ,
Chong‐Yao Jin
1 ,
Nai‐Jia Xue
1 ,
Jia‐Xian Dong
1 ,
Ya‐Ping Yan
1 ,
Jun Tian
1 ,
Jia‐Li Pu
1
,
,
Bao‐Rong Zhang
1
,
14 July 2020
Read this article at
There is no author summary for this article yet. Authors can add summaries to their articles on ScienceOpen to make them more accessible to a non-specialist audience.
Abstract
1
Parkinson's disease (PD) is the second most common neurodegenerative disease after
Alzheimer's disease, affecting approximately 2%‐3% of the global population aged ≥65 years.
1
The loss of dopaminergic (DA) neurons in the substantia nigral pars compacta (SNc)
and intracellular α‐synuclein deposition are known as PD’s pivotal pathological characteristics.
Although its etiology remains to be fully elucidated, the hypothesis that environmental
and genetic factors play essential roles in the initiation and progression of PD is
acknowledged worldwide.
2
Over the past two decades, substantial progress has been made in our understanding
of the genetic basis of parkinsonism through the identification of 19 monogenic disease‐causing
genes. In addition, emerging evidence suggests that some monogenic genes can also
increase the risk of sporadic PD through different mechanisms.
3
Previous studies have demonstrated that mitochondrial dysfunction plays a fundamental
role in the pathogenesis of PD.
4
Moreover, mitochondrial homeostasis has been related to mutations of Parkin, Pink1,
and DJ‐1.
5
Ubiquinol‐cytochrome c reductase core protein I (UQCRC1) is a component of the complex
III in the respiratory chain complex. Recently, Lin et al used a whole‐exome sequencing
and discovered a novel candidate pathogenetic missense variant (c.941A > C p.Y314S)
of the UQCRC1 gene in a Taiwanese family with autosomal dominant parkinsonism.
6
To further investigate the association between UQCRC1 and PD in eastern China, we
performed a UQCRC1 genetic analysis in a cohort of sporadic PD patients and healthy
controls.
We recruited 452 Chinese Han patients with sporadic PD and 450 sex‐, age‐, and ethnicity‐matched
healthy controls from the Second Affiliated Hospital of Zhejiang University (Hangzhou,
China) between October 2016 and January 2019. All study participants were diagnosed
according to the Movement Disorder Society clinical diagnostic criteria for PD.
7
The basic information and demographic characteristics of participants are all shown
in Data S1. All participants provided written informed consent. This study was approved
by the Ethics Committee of the Second Affiliated Hospital of Zhejiang University of
Medicine.
Blood samples were obtained from all participants. Genomic DNA was then extracted
from peripheral blood leukocytes using the QIAamp DNA Blood Mini Kit (Qiagen, Hilden,
Germany) according to the manufacturer's protocol. The exon and intron‐exon boundaries
of the UQCRC1 gene were amplified by polymerase chain reaction. Detailed information
on primers and annealing temperature is shown in Data S2. DNA products were directly
sequenced using a Genetic Analyzer (ABI 3730XL) and analyzed with Chromas software
and Mutation Surveyor software (SoftGenetics).
PD cases and healthy controls were matched for age and sex using a nonparametric test
(Shapiro‐Wilk normality test, P < .05) and Pearson's χ
2 test, respectively. We also used Pearson's χ
2 test, χ
2 with Yates correction test or Fisher exact test to compare the allele frequencies
in PD patients and healthy controls. The Hardy‐Weinberg principle was used to ascertain
the normal distribution of genotype frequencies in PD patients and healthy controls.
Statistical analysis was performed using SPSS20.0 software (SPSS Inc). A two‐tailed
P < .05 was considered as threshold for statistical significance. The pathogenicity
of missense variants was predicted using in silico tools including PolyPhen2, SIFT,
CADD, as recommended by the American College of Medical Genetics and Genomics.
8
In this study, we enrolled 452 sporadic PD cases and 450 healthy controls. There were
no significant differences in age and sex distribution between patients and controls
(Data S1). We identified three synonymous variants (p.V388=, p.A108=, and p.S473=)
and four nonsynonymous variants (p.D215H, p.S4A, p.P267R, and p.N308S) of the UQCRC1
gene. The p.A108 = and p.S4A variants were detected exclusively in PD patients (Table 1).
Moreover, the homozygous p.V388 = variant and the p.P267R variant were identified
separately in two PD patients. However, we did not detect the previously reported
p.Y314S variant (Lin et al, 2019). Analysis using publicly available resources indicated
all four nonsynonymous variants should not be deleterious (Table 2). Furthermore,
the frequencies of the seven variants showed no statistically significant differences
between PD patients and controls. Representative images of the variants are presented
in Data S3.
TABLE 1
Variants identified in UQCRC1 in our study
Chromosomal position
Accession number
Variant
Number of carrying variants
PD vs controls
cDNA
Amino acid
PD
Controls
OR (95%CI)
P
Chr3:48637964
rs140583334
c.1164G>T
p.V388=
42
40
1.050 (0.667‐1.654)
0.833
Chr3:48642187
NA
c.324C>T
p.A108=
1
0
NA
1.000
Chr3:48636585
rs182453765
c.1419C>T
p.S473=
12
14
0.849 (0.388‐1.857)
0.682
Chr3:48641060
rs17080284
c.643G>C
p.D215H
29
31
0.927 (0.549‐1.565)
0.776
Chr3:48647044
NA
c.10T > G
p.S4A
1
0
NA
1.000
Chr3:48638807
rs149245457
c.800C>G
p.P267R
22
15
1.484 (0.759‐2.899)
0.245
Chr3:48638451
rs187641562
c.923A>G
p.N308S
2
3
0.662 (0.110‐3.982)
0.996
Abbreviations: CI, confidence interval; NA, not available; OR, odds ratio; P, P‐value;
PD, Parkinson's disease.
John Wiley & Sons, Ltd
TABLE 2
Allele frequencies and pathogenicity prediction of identified UQCRC1 missense variants
Missense variants
Freq.gnom AD(East Asian)
Freq.ExAC (East Asian)
Freq.1000G (East Asian)
SIFT score
Polyphen2 score
CADD
p.D215H
0.0291
0.0327
0.0238
0.097
0.997
25.4
p.S4A
NA
NA
NA
0.274
0.175
12.69
p.P267R
0.0410
0.0360
0.0526
0.051
0.052
9
p.N308S
0.0034
0.0042
0.002
0.258
0.093
15.41
Note
Threshold values for deleteriousness: SIFT <0.05; polyphen2 >0.86; CADD >15.
Abbreviation: NA, not available.
John Wiley & Sons, Ltd
The UQCRC1 gene encodes ubiquinol‐cytochrome c reductase core protein I, which is
a subunit of complex III of the respiratory chain. As one of eleven subunits of complex
III, UQCRC1 is a nuclear‐encoded protein localized in the inner mitochondrial membrane.
Previous studies of UQCRC1 have focused mainly on its roles in various tumors, such
as malignant pleural mesothelioma and pancreatic ductal adenocarcinoma, suggesting
that UQCRC1 may contribute to carcinogenesis.
9
,
10
Notably, mitochondrial dysfunction also plays a crucial role in the pathogenesis of
PD.
4
,
11
Therefore, it can be hypothesized that UQCRC1 variants may play a role in the development
of PD. Recently, UQCRC1 was identified as a candidate pathogenic gene for PD.
6
Nevertheless, we did not find an association between sporadic PD and UQCRC1 in our
study. Our data do not support that UQCRC1 mutation is a common genetic factor for
sporadic patients in eastern China, but our study cannot rule out its pathogenic role
in PD due to the limited ethnic background and sample size. Furthermore, in consideration
of the role of outer mitochondrial membrane signaling in Parkinson's disease, proteins
residing in or translocating to the outer mitochondrial membrane following mitochondrial
activities deserve closer attention and be important in genetic analyses for PD.
12
In conclusion, our study indicates that UQCRC1 may not be involved in pathogenesis
of PD in eastern China. Further genetic analysis with larger sample sizes from diverse
ethnic populations and functional assays are still needed to clarify the role of UQCRC1
in the pathogenesis of PD.
CONFLICT OF INTEREST
The authors declare no conflicts of interest.
FUNDING INFORMATION
This work was supported by the National Natural Science Foundation of China (No. 81520108010
and 81771216) and the Primary Research and Development Plan of Zhejiang Province (No.
2020C03020).
REFERENCES
1
Poewe
W
,
Seppi
K
,
Tanner
CM
, et al. Parkinson disease. Nat Rev Dis Primers. 2017;3:17013.28332488
2
Pang
SY
,
Ho
PW
,
Liu
HF
, et al. The interplay of aging, genetics and environmental factors in the pathogenesis
of Parkinson's disease. Transl Neurodegener. 2019;8:23.31428316
3
Reed
X
,
Bandres‐Ciga
S
,
Blauwendraat
C
,
Cookson
MR
. The role of monogenic genes in idiopathic Parkinson's disease. Neurobiol Dis. 2019;124:230‐239.30448284
4
Grunewald
A
,
Kumar
KR
,
Sue
CM
. New insights into the complex role of mitochondria in Parkinson's disease. Prog
Neurogibol. 2019;177:73‐93.
5
Larsen
SB
,
Hanss
Z
,
Krüger
R
. The genetic architecture of mitochondrial dysfunction in Parkinson’s disease. Cell
Tissue Res. 2018;373(1):21‐37.29372317
6
Lin
CH
,
Chen
PL
,
Tai
CH
, et al. A clinical and genetic study of early‐onset and familial parkinsonism in
taiwan: an integrated approach combining gene dosage analysis and next‐generation
sequencing. Mov Disord. 2019;34(4):506‐515.30788857
7
Postuma
RB
,
Berg
D
,
Stern
M
, et al. MDS clinical diagnostic criteria for Parkinson's disease. Mov Disord. 2015;30(12):1591‐1601.26474316
8
Richards
S
,
Aziz
N
,
Bale
S
, et al. Standards and guidelines for the interpretation of sequence variants: a joint
consensus recommendation of the American College of Medical Genetics and Genomics
and the Association for Molecular Pathology. Genet Med. 2015;17(5):405‐423.25741868
9
Torricelli
F
,
Saxena
A
,
Nuamah
R
, et al. Genomic analysis in short‐ and long‐term patients with malignant pleura mesothelioma
treated with palliative chemotherapy. Eur J Cancer. 2020;132:104‐111.32339978
10
Wang
Q
,
Li
M
,
Gan
Y
, et al. Mitochondrial protein UQCRC1 is oncogenic and a potential therapeutic target
for pancreatic cancer. Theranostics. 2020;10(5):2141‐2157.32089737
11
Cowan
K
,
Anichtchik
O
,
Luo
S
. Mitochondrial integrity in neurodegeneration. CNS Neurosci Ther. 2019;25(7):825‐836.30746905
12
Lucero
M
,
Suarez
AE
,
Chambers
JW
. Phosphoregulation on mitochondria: Integration of cell and organelle responses.
CNS Neurosci Ther. 2019;25(7):837‐858.31025544
Supporting information
Data S1
Click here for additional data file.
Data S2
Click here for additional data file.
Data S3
Click here for additional data file.
Related collections
Most cited references8
- Record: found
- Abstract: found
- Article: not found
New insights into the complex role of mitochondria in Parkinson’s disease
Carolyn M. Sue, Kishore Kumar, Anne Grünewald (2018)

- Record: found
- Abstract: found
- Article: found
The interplay of aging, genetics and environmental factors in the pathogenesis of Parkinson’s disease
Shirley Pang, Philip Ho, Hui-Fang Liu … (2019)
- Record: found
- Abstract: found
- Article: found
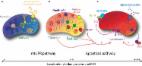
The genetic architecture of mitochondrial dysfunction in Parkinson’s disease
S B Larsen, Z Hanss, R. Kruger (2018)