- Record: found
- Abstract: found
- Article: found
Rhizospheric microbial communities are driven by Panax ginseng at different growth stages and biocontrol bacteria alleviates replanting mortality
Read this article at
Abstract
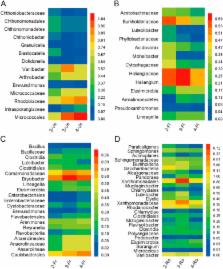
The cultivation of Panax plants is hindered by replanting problems, which may be caused by plant-driven changes in the soil microbial community. Inoculation with microbial antagonists may efficiently alleviate replanting issues. Through high-throughput sequencing, this study revealed that bacterial diversity decreased, whereas fungal diversity increased, in the rhizosphere soils of adult ginseng plants at the root growth stage under different ages. Few microbial community, such as Luteolibacter, Cytophagaceae, Luteibacter, Sphingomonas, Sphingomonadaceae, and Zygomycota, were observed; the relative abundance of microorganisms, namely, Brevundimonas, Enterobacteriaceae, Pandoraea, Cantharellales, Dendryphion, Fusarium, and Chytridiomycota, increased in the soils of adult ginseng plants compared with those in the soils of 2-year-old seedlings. Bacillus subtilis 50-1, a microbial antagonist against the pathogenic Fusarium oxysporum, was isolated through a dual culture technique. These bacteria acted with a biocontrol efficacy of 67.8%. The ginseng death rate and Fusarium abundance decreased by 63.3% and 46.1%, respectively, after inoculation with B. subtilis 50-1. Data revealed that microecological degradation could result from ginseng-driven changes in rhizospheric microbial communities; these changes are associated with the different ages and developmental stages of ginseng plants. Biocontrol using microbial antagonists alleviated the replanting problem.
Graphical abstract
Ginseng cropping induced changes in rhizospheric microbial communities and decreased bacterial diversity. These effects could collectively cause microecological degradation, which consequently results in replanting problems. However, inoculation with a biocontrol bacterial strain alleviated the replanting problem and improved the growth of ginseng. These results provide insight into the reasons that underlie the replanting issues caused by rhizospheric microbial communities.
Related collections
Most cited references48
- Record: found
- Abstract: not found
- Article: not found
GeneMarkS: a self-training method for prediction of gene starts in microbial genomes. Implications for finding sequence motifs in regulatory regions.
- Record: found
- Abstract: found
- Article: not found
Rhizosphere microbiome assemblage is affected by plant development.

- Record: found
- Abstract: found
- Article: found
