- Record: found
- Abstract: found
- Article: not found
Global Genomics and Proteomics Approaches to Identify Host Factors as Targets to Induce Resistance Against Tomato Bushy Stunt Virus
Abstract
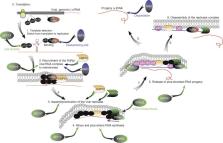
The success of RNA viruses as pathogens of plants, animals, and humans depends on their ability to reprogram the host cell metabolism to support the viral infection cycle and to suppress host defense mechanisms. Plus-strand (+)RNA viruses have limited coding potential necessitating that they co-opt an unknown number of host factors to facilitate their replication in host cells. Global genomics and proteomics approaches performed with Tomato bushy stunt virus (TBSV) and yeast ( Saccharomyces cerevisiae) as a model host have led to the identification of 250 host factors affecting TBSV RNA replication and recombination or bound to the viral replicase, replication proteins, or the viral RNA. The roles of a dozen host factors involved in various steps of the replication process have been validated in yeast as well as a plant host. Altogether, the large number of host factors identified and the great variety of cellular functions performed by these factors indicate the existence of a truly complex interaction between TBSV and the host cell. This review summarizes the advantages of using a simple plant virus and yeast as a model host to advance our understanding of virus–host interactions at the molecular and cellular levels. The knowledge of host factors gained can potentially be used to inhibit virus replication via gene silencing, expression of dominant negative mutants, or design of specific chemical inhibitors leading to novel specific or broad-range resistance and antiviral tools against (+)RNA plant viruses.
Related collections
Most cited references146
- Record: found
- Abstract: found
- Article: not found
Receptor downregulation and multivesicular-body sorting.
- Record: found
- Abstract: found
- Article: not found
RNA interference screen for human genes associated with West Nile virus infection.
- Record: found
- Abstract: found
- Article: not found
